|
Now that we can clearly see everything, we are one step closer to understanding what it all means.” The evolving centromere “Truly finishing the human genome sequence was like putting on a new pair of glasses. “In the future, when someone has their genome sequenced, we will be able to identify all of the variants in their DNA and use that information to better guide their health care,” said Adam Phillippy, one of the leaders of T2T and a senior investigator at the National Human Genome Research Institute (NHGRI) of the National Institutes of Health. They also found about 2 million additional variants in the human genome, 622 of which occur in medically relevant genes. Of the protein-coding genes, the T2T team found about 2,000 new ones, most of them disabled, but 115 of which may still be expressed. The consortium’s gapless version of all 22 autosomes and the X sex chromosome is composed of 3.055 billion base pairs, the units from which chromosomes and our genes are built, and 19,969 protein-coding genes. The sequencing and analysis were performed by a team of more than 100 people, the so-called Telemere-to-Telomere Consortium, or T2T, named for the telomeres that cap the ends of all chromosomes. Four companion papers, including one for which Altemose is co-first author, also will appear online April 1 in the journal Nature Methods. A paper explaining how the sequencing was done will appear in the April 1 print edition of the journal Science, while Altemose’s centromere paper and four others describing what the new sequences tell us are summarized in the journal with the full papers posted online. “Before, we just had the blurriest picture of what was there, and now it’s crystal clear down to single base pair resolution.”Īltemose is first author of one paper that describes the base pair sequences around the centromere. “Uncovering the complete sequence of these formerly missing regions of the genome told us so much about how they’re organized, which was totally unknown for many chromosomes,” said Nicolas Altemose, a postdoctoral researcher at the University of California, Berkeley, and co-author of four new articles describing the completed genome. Variability within this area might potentially provide fresh information about how our ancestors developed in Africa. The new DNA sequences, among other things, provide previously unknown details about the area around the centromere, which is where chromosomes are seized and tugged apart as cells split, ensuring that each “daughter” cell acquires the right amount of chromosomes.  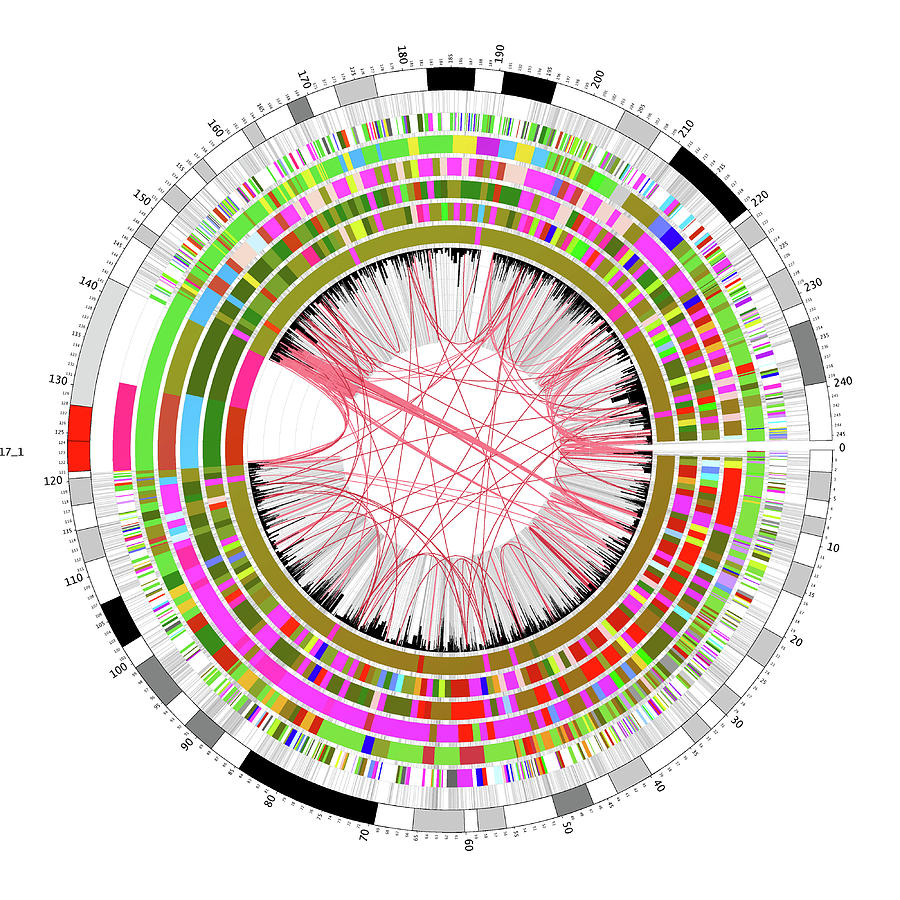
The recently completed genome, termed T2T-CHM13, is a significant improvement over the existing reference genome, GRCh38, which is used by physicians and scientists to check for disease-linked mutations as well as to study the evolution of human genetic diversity. However, a three-year-old team has finally filled in the gaps in the remaining DNA, giving scientists and physicians the first complete, gap-free genome sequencing. 
In actuality, almost 20 years later, approximately 8% of the genome has never been completely sequenced, due to highly repetitive DNA segments that are difficult to match with the rest of the genome. Scientists lied a little when they revealed the entire sequencing of the human genome in 2003. Repetitive DNA sequences around centromere show history of human genetic variation. The entire genome consists of more than 3 billion nucleotides. Sequencing the last 8% of the human genome has taken 20 years and the invention of new techniques for reading long sequences of the genetic code, which consists of the nucleotides C, T, G and A.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
